Genetic Algorithms
Genetic algorithms are complex search algorithms inspired by the theory of evolution. It is a population-based algorithm in which candidate optimisation agents possess genetic information and evolve over time generation after generation through three main evolutionary life processes: elitism, mutation and crossover.

Components of a genetic algorithm
- Genetic Code Formation
- Fitness/Cost Function
- Population Initialisation
- Elitism
- Crossover
- Mutation
When solving a search problem using a genetic algorithm, the genetic code must be defined. The genetic code refers to the nature or profile of the candidate solution. For instance, a search problem of finding the lowest point in a multivariate function will have as genetic code the coordinate vector (i.e. x = [x1,x2,…xn]) of a potential solution. For a search problem of finding the shortest route to navigate a maze, the genetic code will refer to the segment set to exit the maze (x = {s1,s2,s3,…sn}) which forms the genetic profile of every population individual.
Upon determining the genetic profile of population individuals, a cost or fitness function (i.e. f(x)) must be defined that will quantify the performance of any given candidate solution. It completes the mathematical formulation of the problem (i.e. min or max f(x)) to be solved leading to the start of the search process. A population of N individuals randomly profiled based on the genetic code is initialised forming the first population generation. An ep portion of the fittest individuals in the population as per the cost function gains immediate access to the next generation. This process is referred to as “elitism” where the fittest individuals survive.
The remaining (N-epN) individuals for the next generation are obtained through crossover and mutation. During crossover, individuals are allowed to mate, exchanging genetic information, with the hope of generating better-suited offspring. Mating is typically performed between elites or between elites and other individuals in the population. A crossover operator must thus be defined that will specify the mating process’s inherent rules. During mutation, individuals independently modify their own genetic information leading to newer species in a bid to diversity in the search process which may prevent premature convergence to information that is already known. A mutation operator must equally be defined.
This process continues generation after generation until a stop criterion is defined. Most typically, a maximum generation count is used to stop the search process or the search process is heuristically halted when the quality of the solution does not significantly improve over time.

Shortest route search problem - Python implementation
Let’s look at the logistic problem of delivering packages from a main warehouse to several branches across a given region. A genetic algorithm search will be devised to find a good enough navigation path that will minimise transportation costs and time. We assume that all branches have existing physical routes that can interconnect them.
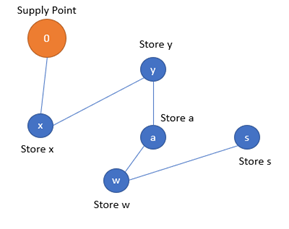
Genetic code formation
The problem consists of finding the best permutation of 15 stores. Thus, the genetic code of a population individual consists of a given permutation of the 15 stores.
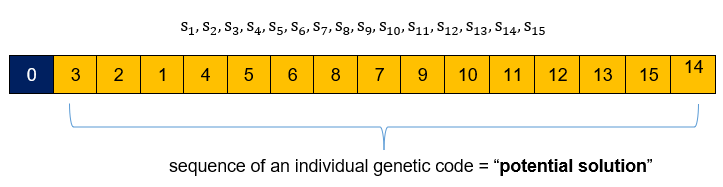
Fitness function
The fitness function to solve the problem is the total distance from the supply point through town to town:
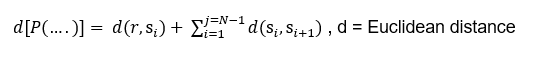
The search problem, thus consist of finding the best permutation will yield the smallest covered distance.
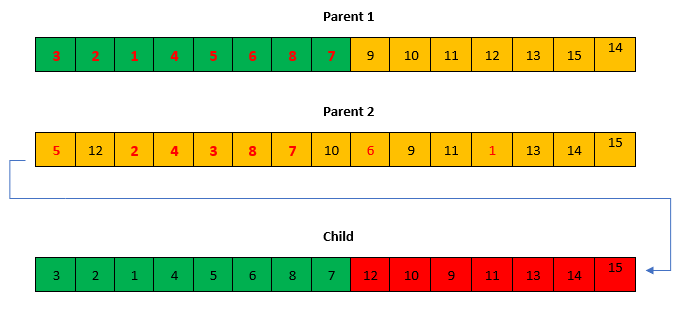
Crossover operator
Crossover is performed by inbreeding two parents to get a better individual of the -two. There could be several inbreeding approaches to the problem, nevertheless, an ordered crossover approach is used in the current implementation. A portion of the genes of one parent is selected and appended in order and no repetition with the remaining genes of the other parent (This process can be used to generate at least two children, parent 2 can be used as parent 1 to generate another child).

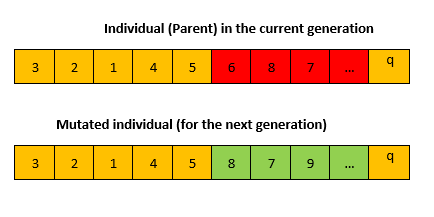
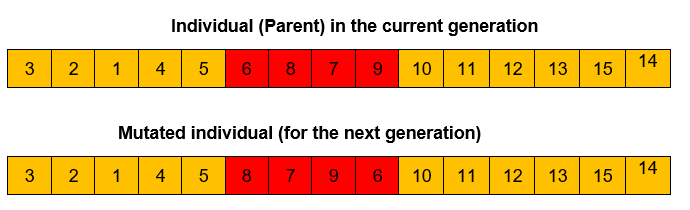
Mutation operator
In the mutation process, a portion of the sequence is selected and reshuffled to create a new individual as described below

Initialisation and settings
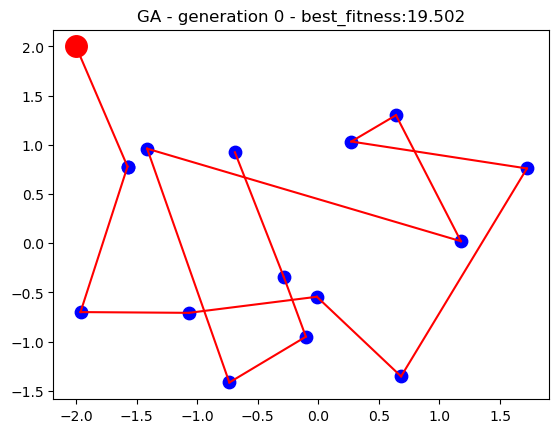
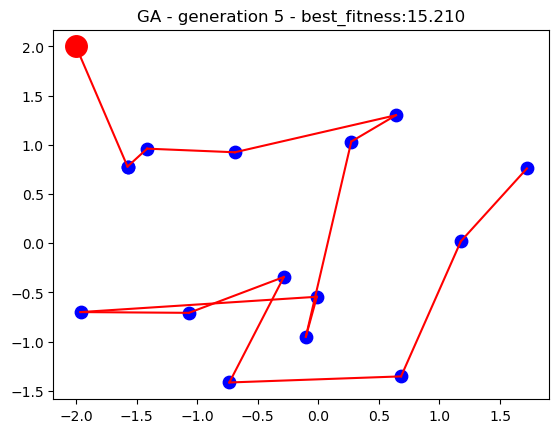
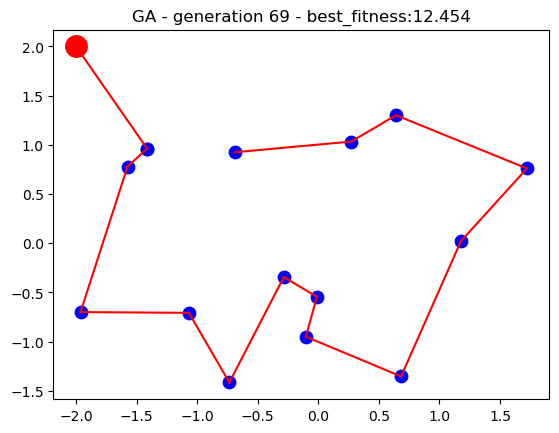
For the current implementation, the location of the main branch and the 15 stores have been loaded from the datasets. A population of 50 search individuals is used for the genetic algorithm and the search runs for up to 50 generations. Upon each generation, the fittest individual through the search is dynamically updated as the best solution thus far until the search process stops. The performance of the algorithm is plotted on a 2D plot over time to visualise how genetic algorithm solutions improve the longer the search runs.
#Import packages
import numpy as np
import matplotlib.pyplot as plt
import copy
#Model stations as objects
class Station:
num_station = 0 #number of stations
def __init__(self,x,y,name):
self.id = Station.num_station
self.x = x
self.y = y
self.name = name
Station.num_station = Station.num_station + 1
def getStationPerId(Stations,id):#return station based on id
numS = len(Stations)
for i in range(0,numS):
mStation = Stations[i]
if(mStation.id == id):
return mStation
#Create plot function for stations
def plotStations(Stations,rootStation,seq,title=''):
node_size = 80
numS = len(Stations)
plt.figure(1)
plt.scatter(rootStation.x,rootStation.y,s=int(3*node_size),c="r")
Station1 = getStationPerId(Stations,seq[0])#link root with first station
X = np.array([Station1.x,rootStation.x])#X array
Y = np.array([Station1.y,rootStation.y])#Y array
plt.plot(X,Y,'r-')#plot line between two stations
plt.scatter(Station1.x,Station1.y,s=node_size,c="b")#plot station 1
for i in range(0,numS-1):
Station1 = getStationPerId(Stations,seq[i])
Station2 = getStationPerId(Stations,seq[i+1])
X = np.array([Station1.x,Station2.x])#X array
Y = np.array([Station1.y,Station2.y])#Y array
plt.plot(X,Y,'r-')#plot line between two stations
plt.scatter(Station1.x,Station1.y,s=node_size,c="b")#plot station 1
Station1 = getStationPerId(Stations,seq[numS-1])
plt.scatter(Station1.x,Station1.y,s=node_size,c="b")#plot station 1
plt.title(title)
plt.show()
#Create objective functions
def dist(x1,y1,x2,y2):
d = np.sqrt((x1-x2)*(x1-x2) + (y1-y2)*(y1-y2) )
return d
def objFunc(Stations,rootStation,seq):#find the accumulated distance between town
numS = len(Stations)
Station1 = getStationPerId(Stations,seq[0])
d = dist(rootStation.x,rootStation.y,Station1.x,Station1.y)
for i in range(0,numS-1):
Station1 = getStationPerId(Stations,seq[i])
Station2 = getStationPerId(Stations,seq[i+1])
d = d+dist(Station2.x,Station2.y,Station1.x,Station1.y)
return d
#Create Crossover Operator
def xover_func(seq1,seq2,ch):#mate the sequence
Ns = len(seq1)
#separate sequence in twos
mi = np.math.floor(Ns/2) #middle index
seq11 = seq1[0:mi]
seq21 = seq2[0:mi]
if ch==1: #get first child
child = seq11
for k in range(0,Ns):
v = np.where(child == seq2[k])
if(len(v[0]) == 0):#element not in sequence
child = np.concatenate((child,[seq2[k]]))
else:
child = seq21
for k in range(0,Ns):
v = np.where(child == seq1[k])
if(len(v[0]) == 0):#element not in sequence
child = np.concatenate((child,[seq1[k]]))
return child
#Create Muation Operator
def mutate_func(seq1,mr):
Ns = len(seq1)#lenght of sequence
Mn = np.math.floor(Ns*mr)#mutation number
dice = np.arange(0,10)
np.random.shuffle(dice)
if dice[0] < 5:
startr = np.arange(0,Ns-Mn)
for i in range(1,10):
np.random.shuffle(startr)
start = startr[0]
seg = copy.deepcopy(seq1[start:(start+Mn)])
np.random.shuffle(seg)
child0 = copy.deepcopy(seq1[0:start])
childn = copy.deepcopy(seq1[(start+Mn):Ns])
child = np.concatenate((child0,seg,childn))#mutated
else:
endr = np.arange(Mn-1,Ns)
for i in range(1,10):
np.random.shuffle(endr)
end = endr[0]
#print("end:"+str(end))
seg = copy.deepcopy(seq1[(end-Mn+1):(end+1)])
np.random.shuffle(seg)
child0 = copy.deepcopy(seq1[0:(end-Mn+1)])
childn = copy.deepcopy(seq1[(end+1):Ns])
child = np.concatenate((child0,seg,childn))#mutated
return child
#Define the genetic process
class Sequence:#class for sequence
def __init__(self,seq):
self.seq = seq
self.obj_val = np.Inf
def ga_operation(gen_limit,n,max_gen):
seq = np.arange(1,n)
BestChild = Sequence(seq)#BestChild
BestChild.obj_val = objFunc(Stations,rootStation,BestChild.seq)
Pop = []
NewGenPop = []
for g in range(0,gen_limit):
print("Generation "+str(g)+"....")
#population generation
if g == 0: #first generation
for i in range(0,max_gen):
np.random.shuffle(seq)
pop = copy.deepcopy(Sequence(seq))
pop.obj_val = objFunc(Stations,rootStation,pop.seq)
Pop.append(pop)#create an initial population
#print("Po_seq: "+str(Pop[i].seq))
else:
Pop = NewGenPop
for i in range(0,max_gen):#evaluate fitness
Pop[i].obj_val = objFunc(Stations,rootStation,Pop[i].seq)
#print("Po_seq: "+str(Pop[i].seq))
#sort population based on objective value
for i in range(0,max_gen):
for j in range(0,MaxGen-1):
if Pop[j].obj_val > Pop[j+1].obj_val:#ascending order
temp = Pop[j]
Pop[j] = Pop[j+1]
Pop[j+1] = temp
#pick the current best
if BestChild.obj_val > Pop[0].obj_val: #the best of the elite is the optimum
BestChild = copy.deepcopy(Pop[0])
print("Sequence: "+str(BestChild.seq))
print("Obj func: "+str(BestChild.obj_val)+"\n")
#plot current status
title = 'GA - generation %d - best_fitness:%.3f'%(g,BestChild.obj_val)
plotStations(Stations,rootStation,BestChild.seq,title)#plot all stations
if (g+1) == gen_limit:
return BestChild #break the loop
#pick parents: elite
bestN = int(max_gen*er)
xoverN = int(max_gen*xr)
ElitePop = Pop[0:bestN] #Pick elite children
NewGenPop = ElitePop #passed to new generation already
#crossover
xrNum = 0
eRng = np.arange(0,bestN)#elite range
while xrNum < xoverN:
#crossovering
np.random.shuffle(eRng)
Pr = eRng[0:2]
#print("crosswover parent id:" + str(Pr))
#print("seqs: "+str(ElitePop[0].seq))
#print("seqs: "+str(ElitePop[1].seq))
Parent1 = ElitePop[Pr[0]] #Parent 1
Parent2 = ElitePop[Pr[1]] #Parent 2
seq1 = Parent1.seq
seq2 = Parent2.seq
#print('Elite parent sequence1: '+str(seq1))
#print('Elite parent sequence2: '+str(seq2))
#two children created
Child1 = copy.deepcopy(Sequence(xover_func(seq1,seq2,1)))
Child2 = copy.deepcopy(Sequence(xover_func(seq1,seq2,2)))
#print('Child1:'+str(Child1.seq))
#print('Child2:'+str(Child2.seq))
NewGenPop.append(Child1)
NewGenPop.append(Child2)
xrNum = xrNum + 2
#mutation
mutN = max_gen - bestN - xoverN
mN = 0
while mN < mutN:
#mutation process
coin = np.arange(1,11)
np.random.shuffle(coin)
Child = Sequence([])
if coin[0] < np.math.floor(mf*10): #elite parent mutation
np.random.shuffle(eRng)
Parent = ElitePop[eRng[0]] #Parent under mutation process
seqt = Parent.seq #pick sequence
#print('Parent seq: '+str(Parent.seq))
else:
rng = np.arange(bestN,max_gen)#other elementts
np.random.shuffle(rng)
Parent = Pop[rng[0]]
seqt = Parent.seq #pick sequence
#print('Parent seq: '+str(Parent.seq))
#mutation
Child.seq = mutate_func(seqt,mr) #mutation sequence
NewGenPop.append(Child)#new child
mN = mN + 1#add to mutation
#load stattions locations
import pandas as pd
df = pd.read_csv('https://raw.githubusercontent.com/mlinsights/freemium/main/datasets/metaheuristics/ga/ga_data.csv')
num_stations = len(df)-1
Stations = [] #load stations location
for i in range(0,num_stations):
name = str(df.iloc[i,0])
x = df.iloc[i,1]
y = df.iloc[i,2]
if i==0:
name = "root"
mStation = Station(x,y,name)
rootStation = mStation
else:
mStation = Station(x,y,name)
Stations.append(mStation)
df.head()
#Run Getting Algorithm
#settings-----------------------------
MaxGen = 1000#generate a thousand elements
br = 0.6#pick the best parents proportion
er = 0.3#30% of population are elite
xr = 0.3#30% of population are cross-over
mf = 0.4 #70% of mutations comes from elite
mr = 0.6#mutation ratio
gen_limit = 100
best_individual = ga_operation(gen_limit,num_stations,MaxGen)
print('Problem solution:',best_individual)
Conclusion
Genetic algorithms are powerful complex search algorithms that can be used for discrete value solutions and continuous value solutions with lots of applications in logistics and several self-learning AI. The proper choice of the crossover operator and mutation operator must be investigated on a problem-to-problem basis. The proper balance of mutation and crossover is required as an incline towards crossover favours exploitation, while an incline towards mutation favours exploration. The degree of usage of these operators affects the performance of GA. GA is commonly used in machine learning for wrapper-based feature selection whereby it helps the search for the optimal feature subset that can lead to the best regression or classification performance.
Be the first to receive notification, when new content is available!
